|
By performing a gene ontology analysis, which annotates the molecular targets and biological processes associated with a group of genes, the researchers found that RAE genes were more likely to be involved in immune adaptability and cellular plasticity than biallelic genes. 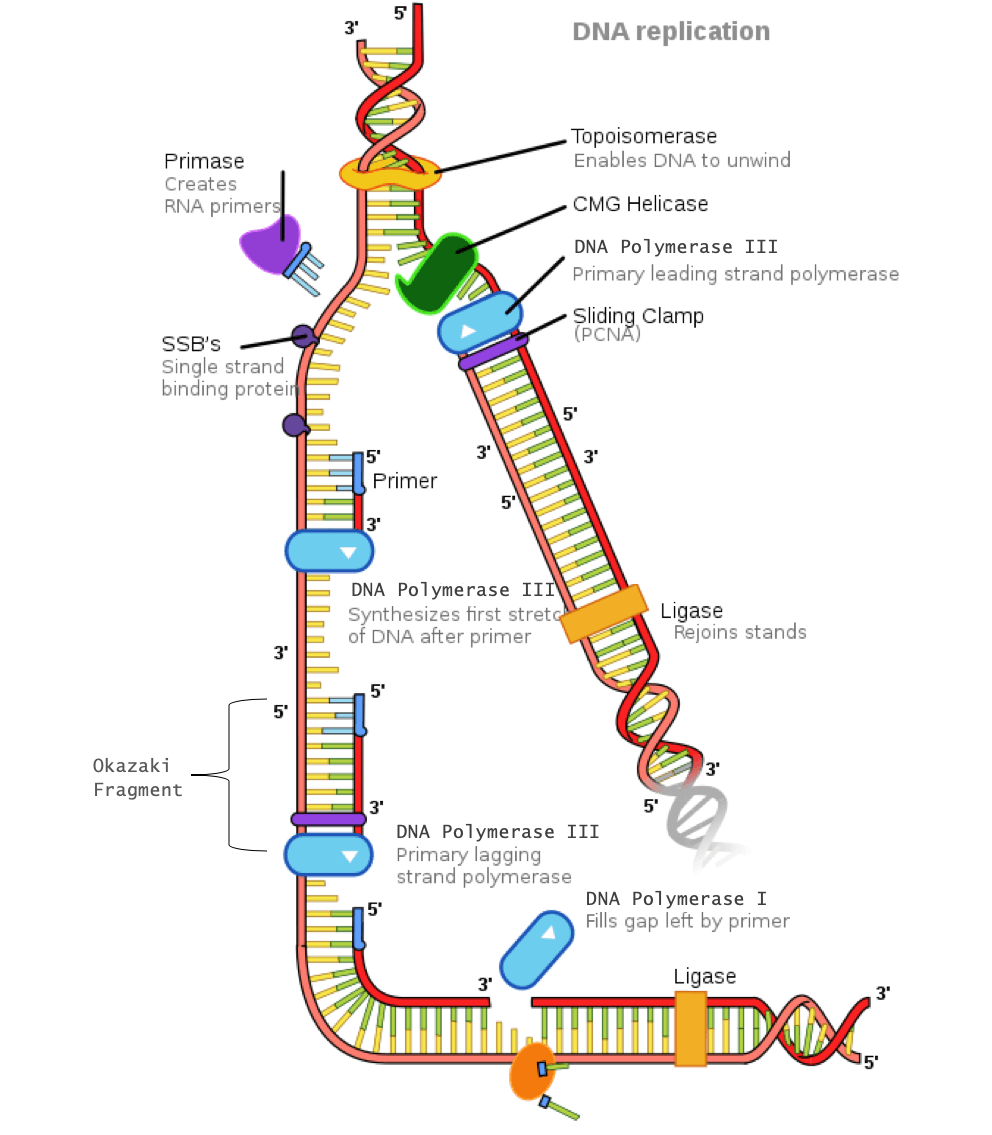
If accurate, that number represents almost 10 percent of known human genes. In all, the analysis picked out 2,762 non–X-linked genes that fit the RAE pattern. As a control, they tested whether the method could distinguish X-linked genes from autosomal genes, and indeed, the model labeled X-linked genes as RAE, validating the model’s ability to discern biased expression of alleles. The researchers then compared the relative gene expression levels of each allele across different tissues within each individual, theorizing that if two alleles are expressed in a biallelic fashion, the overall allelic expression ratio should fit a binomial distribution, whereas randomly expressed alleles wouldn’t. From this data, Kravitz calculated the abundance of each allele across tissues. Wanting to extend the findings to human tissue, he enlisted his former postdoc Stephanie Kravitz to analyze a bulk RNAseq dataset from the genotype tissue expression (GTEx) consortium that included sequences from more than 15,000 genes in 54 tissues from 832 human donors. “But there were other genes that were completely decoupled.” “ discovered that there are some genes where the two alleles are absolutely in lockstep with each other,” Gregg explains. His lab previously found widespread evidence of allelic imbalances in the brains of mice and in cultured cells. Some studies have suggested it occurs in as many as 10 percent of genes, however, Rickard Sandberg, a geneticist at the Karolinska Institute in Sweden, says that his experiments have suggested that imbalanced expression likely occurs in less than one percent of genes.Ĭhristopher Gregg, a neurobiologist and geneticist at the University of Utah school of medicine, suspected that biased allele activity is more common than the literature suggests. And scientists have described cases of random monoallelic expression, wherein cells only express one allele and suppress the other, but experts disagree about the phenomenon’s prevalence. There are some more common exceptions, however, such as how cells with two X chromosomes inactivate one of them during development in order to meter the expression level of X-linked genes that can be toxic in double doses. These genes are more likely to acquire harmful mutations, the authors say, and therefore, could act as biomarkers for predicting or diagnosing disease.īrian Chen, a biologist at McGill University, says the study was “very interesting,” adding: “I definitely thought that the approach was really clever.” Although Chen wasn’t alone in praising the work, the findings sparked some controversy.īiased allele expression is a known genetic phenomenon, but it’s thought to be rare. Using a model that characterizes patterns in allele expression from bulk RNA sequencing data, the researchers found that both alleles of some genes were equally active throughout tissues-but for almost 3,000 genes, cells favored one allele or the other, exhibiting what the authors call random allelic expression (RAE). But a study published in Cell Reports in January posits something different: that it might be somewhat common for cells to preferentially express only one of a gene’s alleles. 
Even though these copies tend to be functionally redundant, conventional genetic theory dictates that biallelic expression, wherein both alleles are transcribed equally when the gene is expressed, is nearly universal. Every person carries two copies of most genes, with one version, or allele, coming from each parent.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
